Web tools of the Laboratory of Chemoinformatics
A tool to color input files. Typing/Coloring means adding information in your input file about different properties found in the molecules.
The different kind of typing are:
• Ph – Pharmacophoric typing
• Ep – Topological electrostatic potentials
• Pc – Partial Charges
• Ff – Force Field coloration
The different kind of typing are:
• Ph – Pharmacophoric typing
• Ep – Topological electrostatic potentials
• Pc – Partial Charges
• Ff – Force Field coloration
Takes SMILES as input and automatically standardizes, types and generates a selected list of ISIDA/Fragments descriptors.
A tool to prepare ligands for the docking tool S4MPLE.
A tool to prepare ligands for the docking tool S4MPLE.
Predictors
Predictors are tools to launch models developed in the lab. They provide predictions as well as confidence scores.
Available predictors:
• CoMet – Conductivity, viscosity, ...
• EU-REACH endpoints – Environmental fate and pathways, ecotoxicological information and human health.
• LogK of imine formation – Regression model predicting the logarithm of equilibrium constant (LogK_eq) of imine formation in chloroform.
• Physprop and activity – Physico-chemical properties and activities predictions.
Available predictors:
• CoMet – Conductivity, viscosity, ...
• EU-REACH endpoints – Environmental fate and pathways, ecotoxicological information and human health.
• LogK of imine formation – Regression model predicting the logarithm of equilibrium constant (LogK_eq) of imine formation in chloroform.
• Physprop and activity – Physico-chemical properties and activities predictions.
The in-house standardization uses the Indigo toolkit.
It consists in the following operations:
- Discarding compounds if the number of atoms is > maxlength
- Normalizing structure using Indigo: neutralizes charges, resolves 5-valence Nitrogen, removes hydrogens
- Dearomatizing: converts molecules/reactions to Kekule form
- Standardizing with the following options:
- Keeping only the largest fragment in the molecule.
- Removing fragments that consist of only a single heavy atom.
- Setting all atoms and bonds to NoStereo.
- Removing all relative stereo groupings.
- Setting all atoms and bonds marked UnknownStereo to NoStereo.
- Setting all atoms marked UnknownStereo to NoStereo.
- Setting all bonds marked UnknownStereo to NoStereo.
- Clearing any atom valence query features and resets all implicit hydrogen counts to their standard values.
- Setting the charges on a molecule to a standard form.
- Applying the following 3 reactions, added in-house :
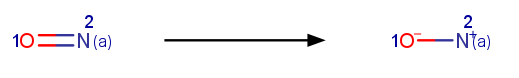

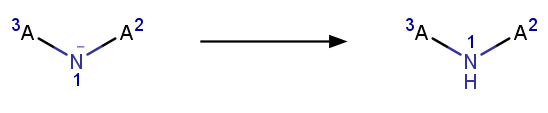
- Aromatizing: Converts molecules/reactions back to aromatic form
- Computing the canonical SMILES (also known as absolute SMILES) string .
